Loading...
Searching...
No Matches
Include dependency graph for init.c:

Go to the source code of this file.
Functions | |
| SEXP | parse_newick (SEXP, SEXP, SEXP) |
| SEXP | newick (SEXP, SEXP) |
| tree in newick format | |
| SEXP | getInfo (SEXP) |
| SEXP | genealSum (SEXP) |
| combine genealogies | |
| SEXP | curtail (SEXP, SEXP, SEXP) |
| curtail the given genealogy | |
| SEXP | yaml (SEXP) |
| extract a YAML description | |
| SEXP | gendat (SEXP, SEXP) |
| data-frame format | |
| SEXP | geneal (SEXP) |
| extract the bare genealogy | |
| DECLARATIONS (LBDP) | |
| DECLARATIONS (Moran) | |
| DECLARATIONS (S2I2R2) | |
| DECLARATIONS (SEIR) | |
| DECLARATIONS (SI2R) | |
| DECLARATIONS (SIIR) | |
| DECLARATIONS (SIR) | |
| DECLARATIONS (Strains) | |
| DECLARATIONS (TwoSpecies) | |
| DECLARATIONS (TwoUndead) | |
| void | R_init_phylopomp (DllInfo *info) |
Variables | |
| get_userdata_t * | get_userdata |
| get_userdata_double_t * | get_userdata_double |
| get_userdata_int_t * | get_userdata_int |
| static const R_CallMethodDef | callMethods [] |
| static const R_CallMethodDef | extMethods [] |
Function Documentation
◆ curtail()
| SEXP curtail | ( | SEXP | State, |
| SEXP | Time, | ||
| SEXP | Troot ) |
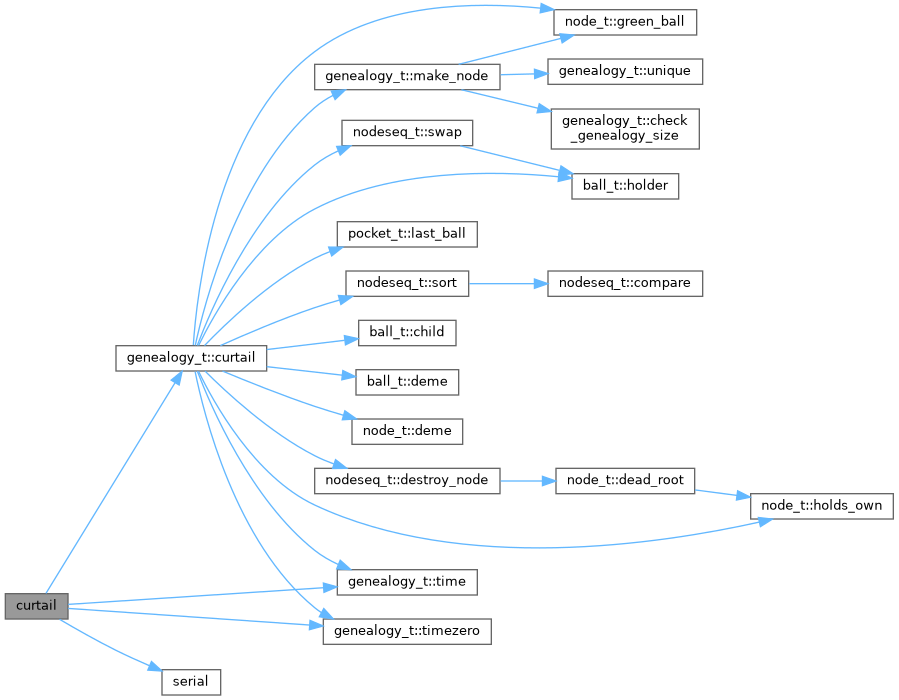
curtail the given genealogy
Definition at line 89 of file curtail.cc.
89 {
90 genealogy_t A = State;
91 double t, t0;
92 t = *REAL(AS_NUMERIC(Time));
93 t0 = *REAL(AS_NUMERIC(Troot));
96 A.curtail(t,t0);
97 SEXP out;
98 PROTECT(out = serial(A));
99 SET_ATTR(out,install("class"),mkString("gpgen"));
100 UNPROTECT(1);
101 return out;
102 }
Here is the call graph for this function:
Here is the caller graph for this function:

◆ DECLARATIONS() [1/10]
| DECLARATIONS | ( | LBDP | ) |
◆ DECLARATIONS() [2/10]
| DECLARATIONS | ( | Moran | ) |
◆ DECLARATIONS() [3/10]
| DECLARATIONS | ( | S2I2R2 | ) |
◆ DECLARATIONS() [4/10]
| DECLARATIONS | ( | SEIR | ) |
◆ DECLARATIONS() [5/10]
| DECLARATIONS | ( | SI2R | ) |
◆ DECLARATIONS() [6/10]
| DECLARATIONS | ( | SIIR | ) |
◆ DECLARATIONS() [7/10]
| DECLARATIONS | ( | SIR | ) |
◆ DECLARATIONS() [8/10]
| DECLARATIONS | ( | Strains | ) |
◆ DECLARATIONS() [9/10]
| DECLARATIONS | ( | TwoSpecies | ) |
◆ DECLARATIONS() [10/10]
| DECLARATIONS | ( | TwoUndead | ) |
◆ gendat()
| SEXP gendat | ( | SEXP | State, |
| SEXP | Obscure ) |
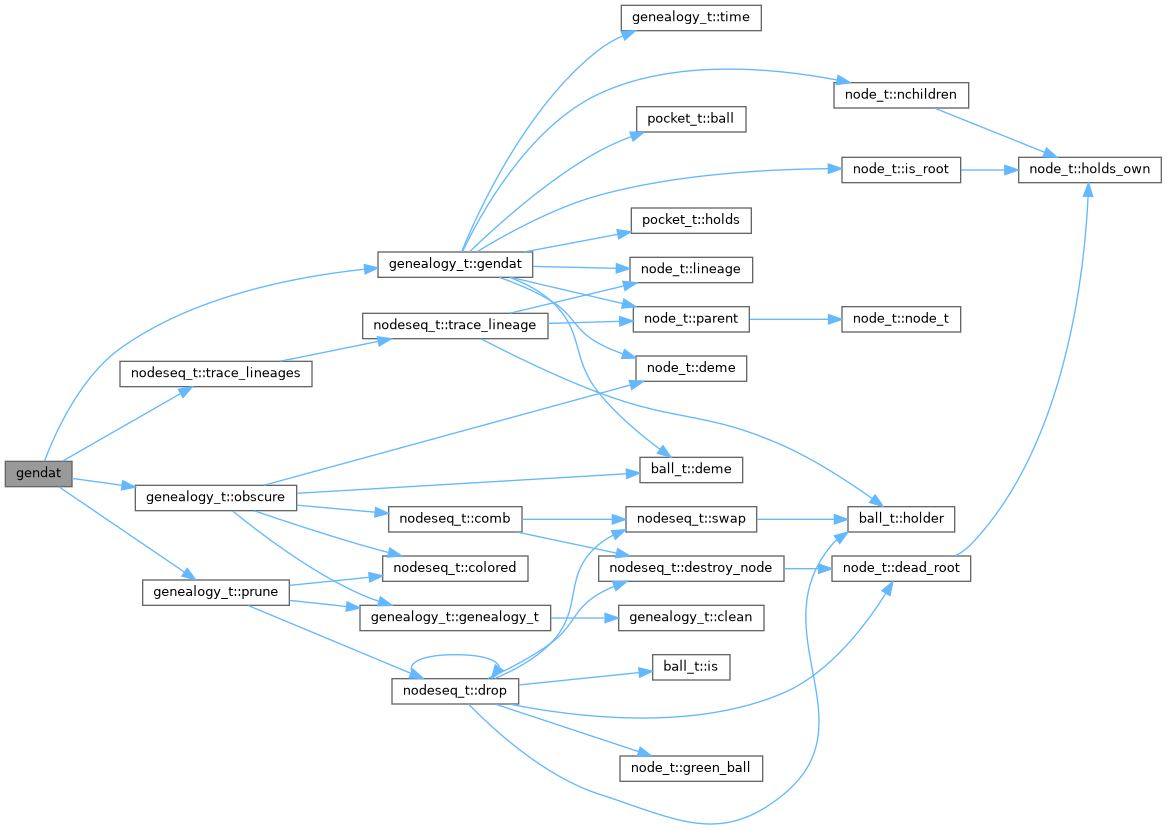
data-frame format
Definition at line 104 of file gendat.cc.
104 {
105 genealogy_t A = State;
106 A.prune();
108 A.trace_lineages();
110 }
void gendat(double *tout, int *anc, int *lin, int *sat, int *type, int *deme, int *index, int *child) const
genealogy information in list format
Definition gendat.cc:9
Here is the call graph for this function:
Here is the caller graph for this function:

◆ geneal()
| SEXP geneal | ( | SEXP | State | ) |
◆ genealSum()
| SEXP genealSum | ( | SEXP | args | ) |
◆ getInfo()
| SEXP getInfo | ( | SEXP | args | ) |
extract requested information prune and/or obscure if requested
Definition at line 19 of file getinfo.cc.
19 {
20 const char *argname[] = {
21 "object","prune","obscure","extended",
22 "t0","time","nsample","nroot","ndeme",
23 "structure","yaml","newick",
24 "lineages","gendat","genealogy"};
25 const int narg = sizeof(argname)/sizeof(const char *);
26 bool flag[narg];
27 SEXP object = R_NilValue;
28 size_t nout = 0;
29 int k;
30
31 for (k = 0; k < narg; k++) flag[k] = false;
32 args = CDR(args);
33
34 while (args != R_NilValue) {
35 const char *name = isNull(TAG(args)) ? "" : CHAR(PRINTNAME(TAG(args)));
36 SEXP arg = CAR(args);
38 if (j == 0) {
39 object = arg;
40 flag[0] = true;
41 } else if (j < 4) {
42 flag[j] = *LOGICAL(AS_LOGICAL(arg));
43 } else if (j < narg) {
44 flag[j] = *LOGICAL(AS_LOGICAL(arg));
45 if (flag[j]) nout++;
46 } else {
48 }
49 args = CDR(args);
50 }
51
53 genealogy_t A = object;
54
55 // prune and/or obscure if requested
56 const bool *f = flag+1;
59 A.trace_lineages();
60 bool extended = false;
61 if (*(f++)) {
62 extended = true;
63 } else {
64 A.insert_zlb();
65 }
66
67 SEXP out, outnames;
68 PROTECT(out = NEW_LIST(nout));
69 PROTECT(outnames = NEW_CHARACTER(nout));
70 k = 0;
71 if (*(f++)) { // t0
73 }
74 if (*(f++)) { // time
76 }
77 if (*(f++)) { // nsample
79 }
80 if (*(f++)) { // nroot
82 }
83 if (*(f++)) { // ndeme
85 }
86 if (*(f++)) { // structure
88 }
89 if (*(f++)) { // yaml
91 }
92 if (*(f++)) { // newick
94 }
95 if (*(f++)) { // lineages
97 }
98 if (*(f++)) { // gendat
100 }
101 if (*(f++)) { // genealogy
102 SEXP S;
106 UNPROTECT(1);
107 }
108 SET_NAMES(out,outnames);
109 UNPROTECT(2);
110 return out;
111 }
static size_t matchargs(const char *prov, const char **set, size_t n)
Definition getinfo.cc:7
static int set_list_elem(SEXP list, SEXP names, SEXP element, const char *name, int pos)
Definition internal.h:76
Here is the call graph for this function:

◆ newick()
| SEXP newick | ( | SEXP | State, |
| SEXP | Extended ) |
tree in newick format
Definition at line 118 of file newick.cc.
118 {
119 PROTECT(Extended = AS_LOGICAL(Extended));
120 bool extended = *LOGICAL(Extended);
121 genealogy_t A(State);
122 UNPROTECT(1);
123 return mkString(A.newick(extended).c_str());
124 }
Here is the call graph for this function:
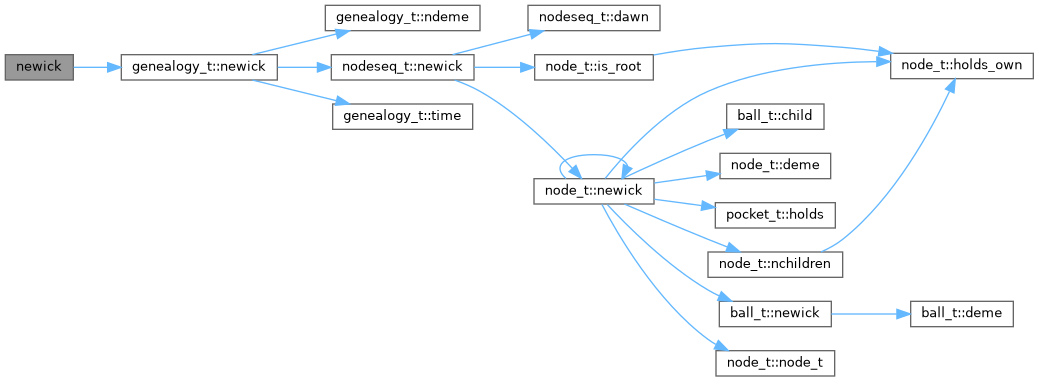
◆ parse_newick()
| SEXP parse_newick | ( | SEXP | X, |
| SEXP | T0, | ||
| SEXP | Tf ) |
A parser for Newick code. Returns a genealogy in the phylopomp format.
Definition at line 259 of file parse.cc.
259 {
260 PROTECT(X = AS_CHARACTER(X));
261 PROTECT(T0 = AS_NUMERIC(T0));
262 PROTECT(Tf = AS_NUMERIC(Tf));
263 double t0 = *REAL(T0);
264 double tf = *REAL(Tf);
265 // parse the Newick representation into a genealogy:
266 string_t x = CHAR(STRING_ELT(X,0));
267 genealogy_t G(t0);
268 G.parse(x);
269 if (!ISNA(tf)) {
270 G.curtail(tf,t0);
271 }
272 G.trace_lineages();
273 UNPROTECT(3);
275 }
Here is the call graph for this function:

◆ R_init_phylopomp()
| void R_init_phylopomp | ( | DllInfo * | info | ) |
◆ yaml()
| SEXP yaml | ( | SEXP | State | ) |
extract a YAML description
Definition at line 78 of file yaml.cc.
78 {
79 genealogy_t A = State;
81 }
Here is the call graph for this function:

Variable Documentation
◆ callMethods
|
static |
Initial value:
= {
METHODS(LBDP),
METHODS(Moran),
METHODS(S2I2R2),
METHODS(SEIR),
METHODS(SI2R),
METHODS(SIIR),
METHODS(SIR),
METHODS(Strains),
METHODS(TwoSpecies),
METHODS(TwoUndead),
{"parse_newick", (DL_FUNC) &parse_newick, 3},
{"newick", (DL_FUNC) &newick, 2},
{"curtail", (DL_FUNC) &curtail, 3},
{"yaml", (DL_FUNC) &yaml, 1},
{"gendat", (DL_FUNC) &gendat, 2},
{"geneal", (DL_FUNC) &geneal, 1},
{NULL, NULL, 0}
}
◆ extMethods
|
static |